TL;DR#
This study explores automating clinical coding in Russian, a language with limited biomedical resources. Existing research focuses on English datasets, leaving a gap in resources for other languages. Assigning ICD codes is crucial for medical documentation but poses challenges due to medical terminology, subjective interpretations, and changing classification standards. Errors in manual coding can lead to misdiagnosis and financial repercussions, highlighting the need for accurate and efficient coding methods.
To address these issues, this paper presents RuCCoD, a new dataset for ICD coding in Russian, labeled by medical professionals. It benchmarks models like BERT and LLaMA and examines transfer learning across domains. Experiments show that training with automatically predicted codes improves accuracy compared to manually annotated data. These findings highlight the potential for automating clinical coding in resource-limited languages and enhancing clinical efficiency and data accuracy.
Key Takeaways#
Why does it matter?#
This paper is important for researchers because it introduces RuCCoD, a novel dataset for ICD coding in Russian, addressing a gap in resources for low-resource languages. It also offers insights into the performance of various models, including LLMs, for automated clinical coding, and demonstrates the potential for improved accuracy in diagnosis prediction using AI-guided coding.
Visual Insights#

🔼 Figure 1 showcases examples from the RuCCoD dataset, illustrating the annotation process for ICD codes. Each highlighted entity (in green) represents a medical concept extracted from a Russian electronic health record (EHR). Above each green entity is its corresponding ICD code, with an English translation provided in yellow. This visualization clarifies how annotators linked specific diagnostic information within EHR text to standard ICD codes.
read the caption
Figure 1: Examples of ICD code assignments by annotators: each entity in green is annotated with its ICD code above and its English translation (in yellow).
| Train | Test | |
|---|---|---|
| # of records | 3000 | 500 |
| # of assigned entities | 8769 | 1557 |
| # of unique ICD codes | 1455 | 548 |
| Avg. # of codes per record | 3 | 3 |
🔼 This table presents a summary of the statistics for the RuCCoD dataset used in the ICD coding task. It shows the number of records, the number of entities assigned ICD codes, the number of unique ICD codes present in the dataset, and the average number of codes per record, broken down for both the training and testing sets of the dataset. This provides a comprehensive overview of the size and complexity of the RuCCoD dataset.
read the caption
Table 1: Statistics for the RuCCoD training and testing sets on ICD coding of diagnosis.
In-depth insights#
ICD Auto-Coding#
ICD auto-coding aims to streamline medical record management by automatically assigning standardized codes to diagnoses. This is critical for billing, insurance, and research. The challenges include navigating complex medical terminology and maintaining accuracy amidst evolving classification standards. Automation can reduce human error and financial repercussions linked to manual coding errors, including financial losses. The process mirrors medical concept normalization, linking physician diagnoses to ICD codes via information extraction. Recent research focuses on neural networks to improve accuracy, but challenges remain, such as data scarcity in languages other than English, variability in clinical notes, and hierarchical ICD code structures. Advanced techniques like BERT, LLMs with PEFT, and RAG are being explored to tackle these hurdles. Further, automatic ICD coding requires integration of external medical knowledge sources like knowledge graphs which will facilitate LLMs generalization on rare codes.
RuCCoD Dataset#
The RuCCoD dataset appears to be a novel resource for ICD coding in Russian, a low-resource language in the biomedical domain. It’s a significant contribution because most existing ICD coding datasets are in English, hindering research and development in other languages. The dataset’s creation aims to address the limitations of using UMLS for ICD coding in Russian, as UMLS may not fully capture the structured semantic requirements of ICD. RuCCoD’s manual annotation by medical professionals, focusing on ICD-10 CM concepts, ensures high-quality labels tailored to clinical practice. The dataset’s size, with over 3,000 records, and the annotation process emphasizing inter-annotator agreement, indicates a commitment to reliability. The use of RuCCoD as a benchmark for state-of-the-art models and its application in downstream tasks highlights its potential for advancing automated clinical coding in Russian and bridging the resource gap.
BERT vs LLAMA#
BERT excels in nuanced language understanding through its bidirectional training, capturing contextual relationships effectively. It’s computationally intensive but yields high accuracy in various NLP tasks. LLaMA, a large language model, prioritizes efficient inference and generation. While it may not match BERT’s contextual depth, its speed and scalability make it suitable for real-time applications. The choice hinges on balancing accuracy needs with resource constraints.
EHR Improvement#
EHR improvement through AI, as discussed in the paper, centers on enhancing data accuracy and utility. The study shows auto-labeling EHRs improves diagnostic model performance, surpassing doctor-assigned codes. This highlights AI’s potential to refine data entry, reduce errors, and standardize clinical info. Key is using AI to pre-train models on vast, auto-labeled datasets, aiding disease diagnosis. The research underscores AI’s role in advancing data quality for better healthcare outcomes and efficiency.
AI-Driven Assist.#
AI-driven assistance in medical coding and diagnosis holds immense potential. By automating ICD coding, AI can reduce errors, enhance clinical efficiency, and improve data accuracy, especially in resource-limited languages. AI can provide independent opinions potentially beneficial in decision-making. Automating and providing better outcomes than manual data annotation by physicians proves the complexity of the ICD system for doctors and shows that AI can assist doctors in the diagnosis. AI helps address challenges like terminology navigation and classification standard updates. Furthermore, automating enables comprehensive analysis, thus aiding early disease identification.
More visual insights#
More on figures

🔼 This figure illustrates the two main tasks addressed in the paper: ICD coding and diagnosis prediction. The ICD coding task (shown in blue) involves using a doctor’s diagnostic conclusion (from the current visit) to assign ICD codes. An AI model is trained to perform this task, generating AI-assigned ICD codes. The diagnosis prediction task (shown in yellow) predicts likely ICD codes based on a patient’s complete medical history, excluding the doctor’s conclusion from the current visit. Both the original ICD codes (assigned by doctors) and the AI-generated ICD codes are used as training targets for the diagnosis prediction models, enabling comparison and improvement of AI performance.
read the caption
Figure 2: Schematic description of ICD coding (in blue) and diagnosis prediction tasks (in yellow). Diagnosis prediction uses prior EHR data and current visit details, excluding the doctor’s conclusion, which is used for ICD coding to generate AI-assigned ICD codes. Both original and AI ICD code lists are then used as targets to train different diagnosis prediction models.
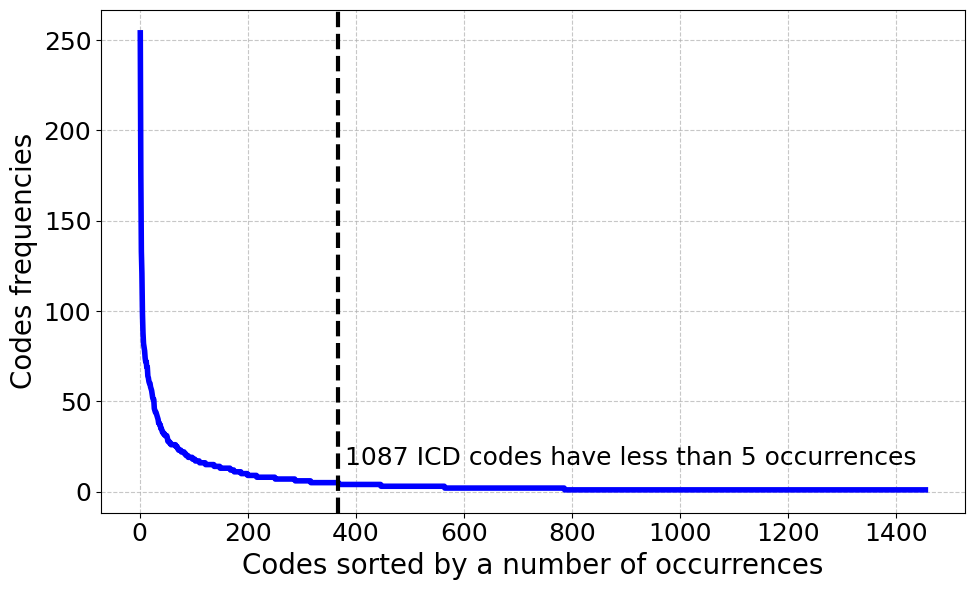
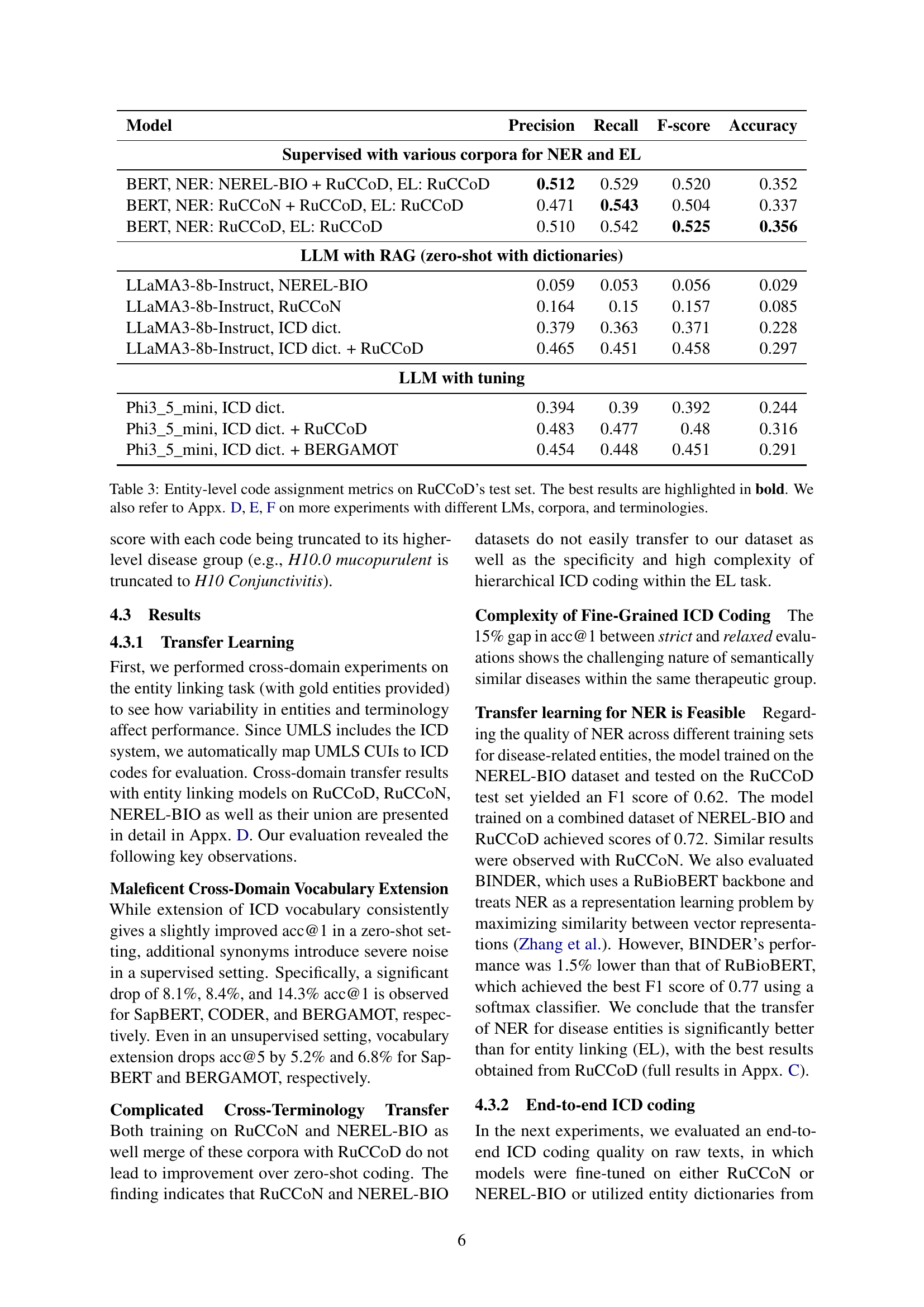
🔼 This histogram displays the frequency distribution of ICD codes within the RuCCoD training dataset. The x-axis represents ICD codes sorted by their frequency of appearance in the dataset, and the y-axis shows the number of times each code appears. The graph reveals a highly skewed distribution, with a relatively small number of codes appearing very frequently, and a large number of codes appearing very infrequently, reflecting the uneven distribution of diagnoses in real-world clinical data.
read the caption
Figure 3: Distribution of ICD code frequencies in the RuCCoD train set.
🔼 This figure shows the performance comparison between two models trained for diagnosis prediction on a manual test set. One model was trained using the original dataset (manually annotated), while the other was trained using a linked dataset (automatically annotated using ICD codes generated by a model). The x-axis represents the training steps, and the y-axis represents the weighted F1-score, a metric that accounts for class imbalances in the dataset. The graph illustrates how the weighted F1-score changes as the models are trained over different numbers of steps. It shows that the model trained on the automatically annotated dataset performs significantly better than the model trained on the original dataset.
read the caption
Figure 4: Comparison of weighted F1 scores on the manual diagnosis prediction test set for models trained on original and linked datasets at different training steps.
🔼 This figure displays the F1-score distribution for the top and bottom 10% most frequent ICD codes within a common test set, comparing performance of the model trained on the original dataset versus the linked dataset. The top 10% represents the most frequently occurring ICD codes, while the bottom 10% represents the least frequent codes, with a minimum frequency threshold of 15 instances within the test set. This visualization highlights the effect of training data (original vs. linked) on the model’s ability to predict both frequent and infrequent disease codes.
read the caption
Figure 5: F1 score distribution for top and bottom 10% frequent ICD codes in the common test set.
More on tables
| Original Dataset | Linked Dataset | Manual Test Set | |
| Number of records | 865539 | 865539 | 494 |
| Number of unique patients | 164527 | 164527 | 450 |
| Number of unique ICD codes | 3546 | 3546 | 394 |
| Avg. number of ICD codes per patient | |||
| Avg. number of EHR records before current appointment | (15, 36, 73) | (15, 36, 73) | (17, 36, 77) |
| Avg. length of EHR records per one appointment | (77, 167, 316) | (77, 167, 316) | (86, 176, 320) |
| Patient’s age | (59, 67, 74) | (59, 67, 74) | (60, 67, 75) |
| Percentage of male patients | 69 | 69 | 71 |
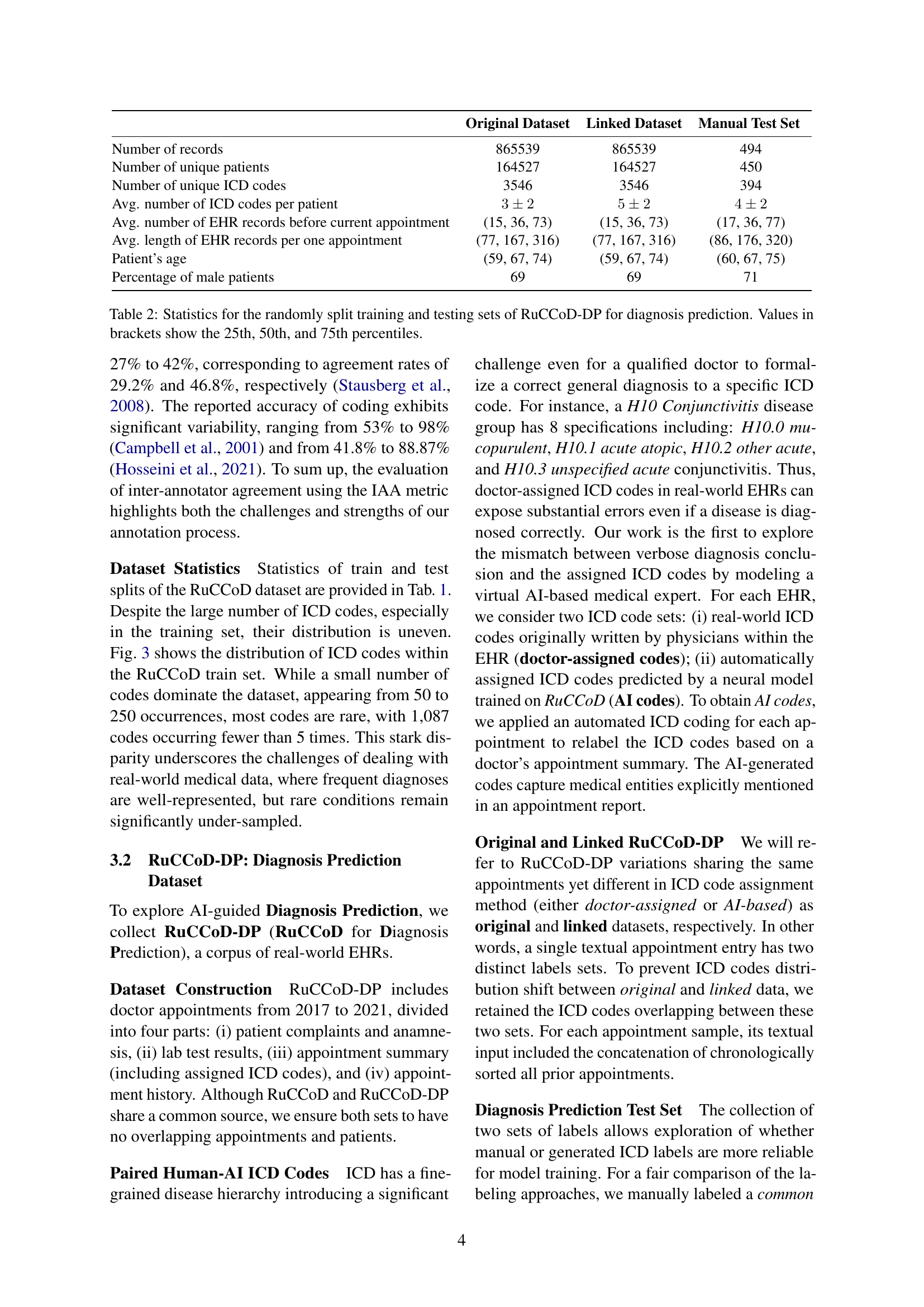
🔼 This table presents a statistical overview of the RuCCoD-DP dataset, which is used for diagnosis prediction. RuCCoD-DP is a collection of real-world electronic health records (EHRs) divided into training and testing sets. The table shows the number of records, unique patients, and unique ICD codes in each set. It also provides the average number of ICD codes per patient, the average number of EHR records per patient before the current appointment, the average length of EHR records per appointment, the average age of patients, and the percentage of male patients. The values in parentheses represent the 25th, 50th, and 75th percentiles, giving a clearer picture of the data distribution.
read the caption
Table 2: Statistics for the randomly split training and testing sets of RuCCoD-DP for diagnosis prediction. Values in brackets show the 25th, 50th, and 75th percentiles.
| Model | Precision | Recall | F-score | Accuracy |
|---|---|---|---|---|
| Supervised with various corpora for NER and EL | ||||
| BERT, NER: NEREL-BIO + RuCCoD, EL: RuCCoD | 0.512 | 0.529 | 0.520 | 0.352 |
| BERT, NER: RuCCoN + RuCCoD, EL: RuCCoD | 0.471 | 0.543 | 0.504 | 0.337 |
| BERT, NER: RuCCoD, EL: RuCCoD | 0.510 | 0.542 | 0.525 | 0.356 |
| LLM with RAG (zero-shot with dictionaries) | ||||
| LLaMA3-8b-Instruct, NEREL-BIO | 0.059 | 0.053 | 0.056 | 0.029 |
| LLaMA3-8b-Instruct, RuCCoN | 0.164 | 0.15 | 0.157 | 0.085 |
| LLaMA3-8b-Instruct, ICD dict. | 0.379 | 0.363 | 0.371 | 0.228 |
| LLaMA3-8b-Instruct, ICD dict. + RuCCoD | 0.465 | 0.451 | 0.458 | 0.297 |
| LLM with tuning | ||||
| Phi3_5_mini, ICD dict. | 0.394 | 0.39 | 0.392 | 0.244 |
| Phi3_5_mini, ICD dict. + RuCCoD | 0.483 | 0.477 | 0.48 | 0.316 |
| Phi3_5_mini, ICD dict. + BERGAMOT | 0.454 | 0.448 | 0.451 | 0.291 |
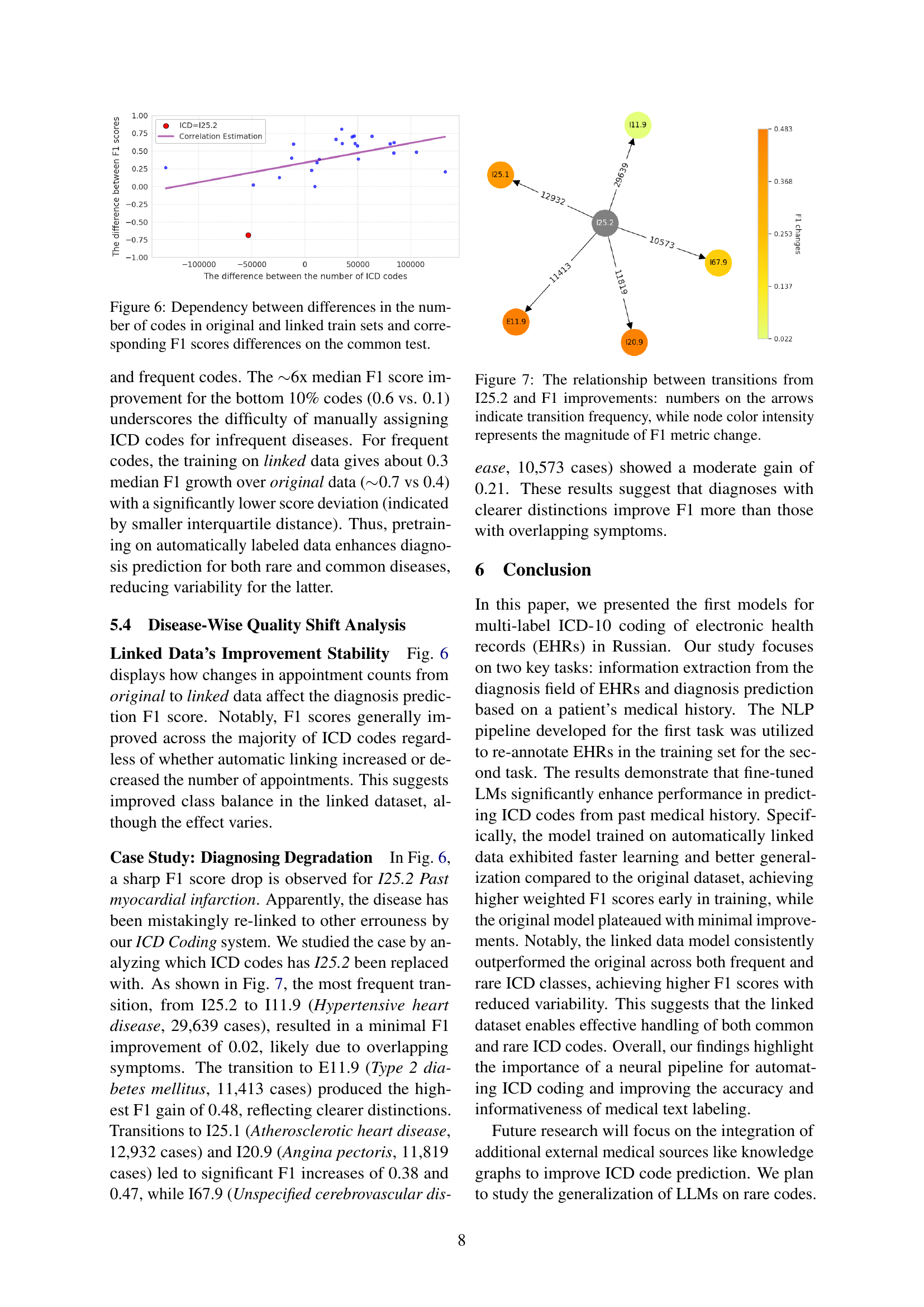
🔼 This table presents the performance of different models on the entity-level code assignment task using the RuCCoD test set. The models evaluated include a BERT-based pipeline, LLMs with PEFT, and LLMs with RAG. The metrics reported are precision, recall, F1-score, and accuracy, allowing for a comprehensive comparison of model effectiveness. The ‘best’ performing model for each metric is highlighted in bold. Further experimental results using different LLMs, corpora, and terminologies are detailed in the Appendix (sections D, E, and F).
read the caption
Table 3: Entity-level code assignment metrics on RuCCoD’s test set. The best results are highlighted in bold. We also refer to Appx. D, E, F on more experiments with different LMs, corpora, and terminologies.
| Task | Model or Approach | LR | # Epochs | BS | Scheduler | WD |
|---|---|---|---|---|---|---|
| NER | RuBioBERT | 1e-5 | 20 | 32 | Cosine Loshchilov and Hutter (2017) | 0.01 |
| EL | BERGAMOT+BioSyn | 2e-5 | 20 | 32 | Adam (Kingma and Ba, 2015) | 0.01 |
| LLM tuning | LoRA | 5e-5 | 33 | 2 | Linear with Warmup | 0.01 |
| ICD code prediction | Longformer | 5e-5 | 2 | 4 | Linear with Warmup | 0.01 |
🔼 This table details the models and training hyperparameters used in the experiments. It lists the learning rate (LR), number of epochs, batch size (BS), scheduler type used for adjusting learning rate during training, and weight decay (WD) values for each model. The models are categorized by task (NER, EL, and ICD code prediction). Understanding these hyperparameters is crucial for replicating and interpreting the experimental results.
read the caption
Table 4: Models and training hyperparameters. LR stands for learning rate, BS for batch size, WD for weight decay
| Model | Train Data | F1-score | Precision | Recall |
|---|---|---|---|---|
| RuBioBERT | RuCCoD train | 0.756 | 0.75 | 0.77 |
| RuBioBERT | BIO-NNE train | 0.62 | 0.57 | 0.67 |
| RuBioBERT | RuCCoD + BioNNE train | 0.72 | 0.75 | 0.70 |
| BINDER + RuBioBERT | RuCCoD train | 0.71 | 0.72 | 0.71 |
🔼 This table presents the performance of different models on the Named Entity Recognition (NER) task using the RuCCoD dataset. It shows the F1-score, precision, and recall achieved by various models trained on different combinations of training data. The models evaluated include RuBioBERT (with and without BINDER), highlighting their effectiveness in extracting relevant entities from clinical texts in the Russian language.
read the caption
Table 5: Evaluation results for NER task on RuCCoD dataset.
| Train set | SapBERT | CODER | BERGAMOT | |||
|---|---|---|---|---|---|---|
| @1 | @5 | @1 | @5 | @1 | @5 | |
| Zero-shot evaluation, strict | ||||||
| ICD dict | 0.3327 | 0.5712 | 0.2631 | 0.4687 | 0.3495 | 0.6170 |
| ICD dict+UMLS synonyms | 0.3546 | 0.5197 | 0.3237 | 0.4765 | 0.3559 | 0.5487 |
| Supervised evaluation, strict | ||||||
| ICD | 0.6132 | 0.8182 | 0.6202 | 0.8169 | 0.6415 | 0.8459 |
| ICD+UMLS sumonyms | 0.5326 | 0.7382 | 0.5358 | 0.7318 | 0.4984 | 0.7253 |
| RuCCoN | 0.3591 | 0.5345 | 0.3598 | 0.5732 | 0.3643 | 0.5313 |
| RuCCoN+ICD | 0.3952 | 0.5732 | 0.3888 | 0.6570 | 0.3817 | 0.5983 |
| NEREL-BIO | 0.3443 | 0.4913 | 0.3378 | 0.5274 | 0.3353 | 0.5113 |
| NEREL-BIO+ICD | 0.3804 | 0.5596 | 0.3804 | 0.6325 | 0.3598 | 0.5525 |
| Zero-shot evaluation, relaxed | ||||||
| ICD dict | 0.4842 | 0.6886 | 0.3752 | 0.6190 | 0.5035 | 0.7286 |
| ICD dict+UMLS synonyms | 0.5551 | 0.6867 | 0.5055 | 0.6293 | 0.5603 | 0.7073 |
| Supervised evaluation, relaxed | ||||||
| ICD | 0.7763 | 0.8839 | 0.7872 | 0.8743 | 0.7917 | 0.8943 |
| ICD+UMLS sumonyms | 0.7788 | 0.8616 | 0.7714 | 0.8860 | 0.7449 | 0.8738 |
| RuCCoN | 0.5235 | 0.6531 | 0.5429 | 0.7208 | 0.5132 | 0.6564 |
| RuCCoN+ICD | 0.5493 | 0.6602 | 0.5770 | 0.7485 | 0.5571 | 0.6873 |
| NEREL-BIO | 0.4803 | 0.6067 | 0.4958 | 0.6634 | 0.4778 | 0.6170 |
| NEREL-BIO+ICD | 0.5455 | 0.6447 | 0.5474 | 0.7292 | 0.5384 | 0.6505 |
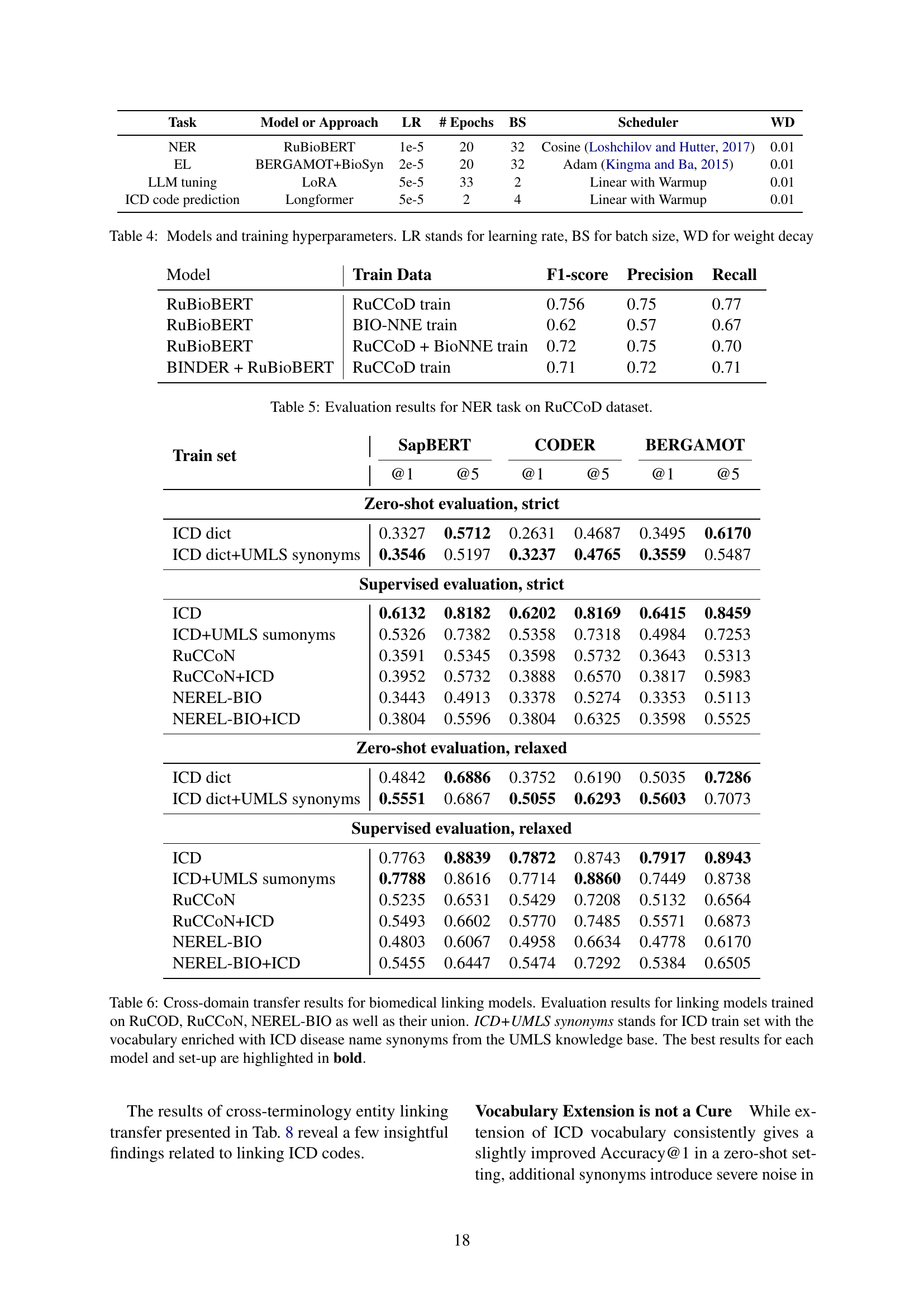
🔼 This table presents the results of experiments evaluating the effectiveness of transfer learning in biomedical entity linking. Four different models (SapBERT, CODER, BERGAMOT) were trained using various combinations of datasets: RuCCoD (Russian ICD Coding Dataset), RuCCON, and NEREL-BIO. Each model was tested with two evaluation methods: ‘strict’ (exact match between predicted and ground truth codes) and ‘relaxed’ (truncated codes to higher level). The results show the precision, recall, F1-score, and accuracy for each model and dataset combination under both strict and relaxed evaluation schemes. One data setting, ICD+UMLS synonyms, involves enriching the training data with disease name synonyms from the UMLS knowledge base to assess the impact of vocabulary expansion.
read the caption
Table 6: Cross-domain transfer results for biomedical linking models. Evaluation results for linking models trained on RuCOD, RuCCoN, NEREL-BIO as well as their union. ICD+UMLS synonyms stands for ICD train set with the vocabulary enriched with ICD disease name synonyms from the UMLS knowledge base. The best results for each model and set-up are highlighted in bold.
| Model | Precision | Recall | F-score | Accuracy |
| NER | ||||
| Llama3-Med42-8B, RuCCoD | 0.642 | 0.642 | 0.642 | 0.473 |
| Qwen2.5-7B-Instruct, RuCCoD | 0.567 | 0.562 | 0.565 | 0.393 |
| Phi3_5_mini, RuCCoD | 0.632 | 0.623 | 0.627 | 0.457 |
| Mistral-Nemo, RuCCoD | 0.631 | 0.598 | 0.614 | 0.443 |
| NER+Linking | ||||
| Llama3-Med42-8B, ICD dict. | 0.149 | 0.149 | 0.149 | 0.08 |
| Llama3-Med42-8B, ICD dict. + RuCCoD | 0.299 | 0.299 | 0.299 | 0.176 |
| Llama3-Med42-8B, ICD dict. + BERGAMOT | 0.286 | 0.286 | 0.286 | 0.167 |
| Qwen2.5-7B-Instruct, ICD dict. | 0.188 | 0.186 | 0.187 | 0.103 |
| Qwen2.5-7B-Instruct, ICD dict. + RuCCoD | 0.281 | 0.279 | 0.28 | 0.163 |
| Qwen2.5-7B-Instruct, ICD dict. + BERGAMOT | 0.2 | 0.198 | 0.199 | 0.11 |
| Phi3_5_mini, ICD dict. | 0.272 | 0.268 | 0.27 | 0.156 |
| Phi3_5_mini, ICD dict. + RuCCoD | 0.335 | 0.33 | 0.333 | 0.199 |
| Phi3_5_mini, ICD dict. + BERGAMOT | 0.322 | 0.317 | 0.32 | 0.19 |
| Mistral-Nemo, ICD dict. | 0.231 | 0.219 | 0.224 | 0.126 |
| Mistral-Nemo, ICD dict. + RuCCoD | 0.303 | 0.287 | 0.295 | 0.173 |
| Mistral-Nemo, ICD dict. + BERGAMOT | 0.267 | 0.253 | 0.26 | 0.149 |
| Code assignment | ||||
| Llama3-Med42-8B, ICD dict. | 0.229 | 0.231 | 0.23 | 0.13 |
| Llama3-Med42-8B, ICD dict. + RuCCoD | 0.434 | 0.435 | 0.435 | 0.278 |
| Llama3-Med42-8B, ICD dict. + BERGAMOT | 0.403 | 0.405 | 0.404 | 0.253 |
| Qwen2.5-7B-Instruct, ICD dict. | 0.296 | 0.295 | 0.295 | 0.173 |
| Qwen2.5-7B-Instruct, ICD dict. + RuCCoD | 0.456 | 0.449 | 0.452 | 0.292 |
| Qwen2.5-7B-Instruct, ICD dict. + BERGAMOT | 0.305 | 0.303 | 0.304 | 0.179 |
| Phi3_5_mini, ICD dict. | 0.394 | 0.39 | 0.392 | 0.244 |
| Phi3_5_mini, ICD dict. + RuCCoD | 0.483 | 0.477 | 0.48 | 0.316 |
| Phi3_5_mini, ICD dict. + BERGAMOT | 0.454 | 0.448 | 0.451 | 0.291 |
| Mistral-Nemo, ICD dict. | 0.326 | 0.311 | 0.319 | 0.189 |
| Mistral-Nemo, ICD dict. + RuCCoD | 0.458 | 0.435 | 0.446 | 0.287 |
| Mistral-Nemo, ICD dict. + BERGAMOT | 0.394 | 0.372 | 0.383 | 0.237 |
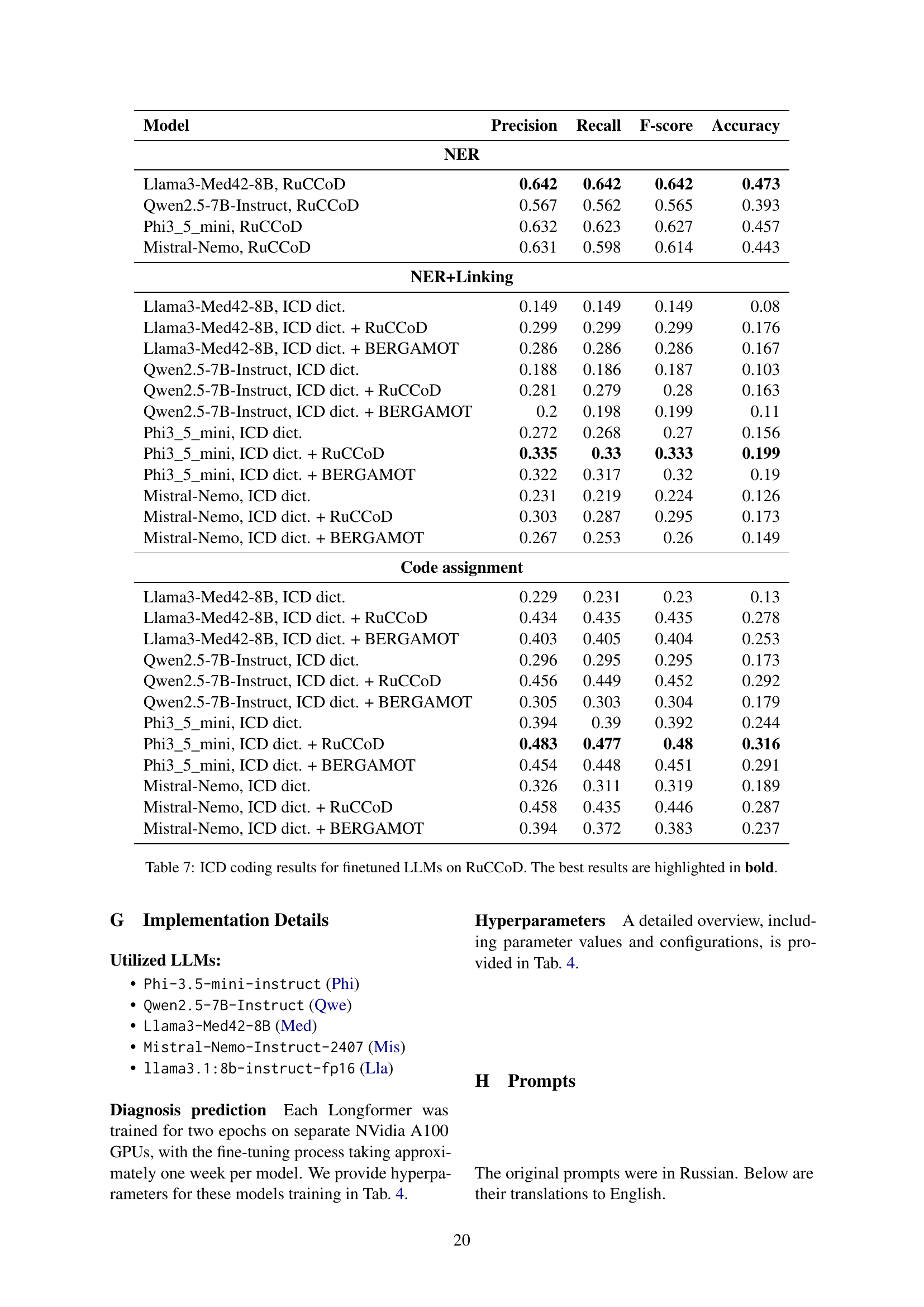
🔼 This table presents the performance of several fine-tuned large language models (LLMs) on the RuCCoD dataset for the task of ICD (International Classification of Diseases) coding. The models were evaluated using micro-averaged precision, recall, F1-score, and accuracy. The table shows the results broken down by the model used and the different corpora employed during training (ICD dict., ICD dict.+RuCCoD, ICD dict.+BERGAMOT). The best-performing model and corpus combination for each metric are highlighted in bold, allowing for a direct comparison across various LLMs and training data configurations.
read the caption
Table 7: ICD coding results for finetuned LLMs on RuCCoD. The best results are highlighted in bold.
| Model | Precision | Recall | F-score | Accuracy |
|---|---|---|---|---|
| NER | ||||
| BioBERT, Biosyn, RuCCoD | 0.649 | 0.655 | 0.653 | 0.485 |
| BioBERT, RuCCoD | 0.721 | 0.769 | 0.744 | 0.592 |
| BioBERT, NEREL-BIO | 0.588 | 0.675 | 0.628 | 0.458 |
| BioBERT, NEREL-BIO, RuCCoD | 0.689 | 0.713 | 0.701 | 0.54 |
| BioBERT, RuCCoN | 0.637 | 0.613 | 0.625 | 0.454 |
| BioBERT, RuCCoN + RuCCoD | 0.609 | 0.709 | 0.655 | 0.487 |
| NER+Linking | ||||
| BioBERT, Biosyn, RuCCoD | 0.392 | 0.396 | 0.394 | 0.245 |
| BioBERT, RuCCoD | 0.427 | 0.455 | 0.441 | 0.283 |
| BioBERT, NEREL-BIO | 0.353 | 0.406 | 0.377 | 0.233 |
| BioBERT, NEREL-BIO, RuCCoD | 0.406 | 0.42 | 0.413 | 0.26 |
| BioBERT, RuCCoN | 0.387 | 0.372 | 0.379 | 0.234 |
| BioBERT, RuCCoN + RuCCoD | 0.351 | 0.409 | 0.378 | 0.233 |
| Code assignment | ||||
| BioBERT, Biosyn, RuCCoD | 0.507 | 0.508 | 0.507 | 0.340 |
| BioBERT, RuCCoD | 0.51 | 0.542 | 0.525 | 0.356 |
| BioBERT, NEREL-BIO | 0.466 | 0.531 | 0.497 | 0.33 |
| BioBERT, NEREL-BIO, RuCCoD | 0.512 | 0.529 | 0.52 | 0.352 |
| BioBERT, RuCCoN | 0.508 | 0.485 | 0.496 | 0.33 |
| BioBERT, RuCCoN + RuCCoD | 0.471 | 0.543 | 0.504 | 0.337 |
🔼 This table presents the performance of a BERT-based information extraction (IE) pipeline on the RuCCoD corpus for three entity-level tasks: Named Entity Recognition (NER), NER + Entity Linking, and ICD Code Assignment. The pipeline uses various combinations of pre-trained models and training corpora (RuCCoD, NEREL-BIO, RuCCON, and BioSyn) for NER and entity linking. The results are shown as precision, recall, F1-score, and accuracy metrics, highlighting the best-performing configurations for each task. This table demonstrates the impact of different model choices and training data on the accuracy of extracting and linking disease-related entities to ICD codes.
read the caption
Table 8: Evaluation results for entity-level tasks for BERT-based IE pipeline on RuCCoD corpus. The best results are highlighted in bold.
| Model | Precision | Recall | F-score | Accuracy |
|---|---|---|---|---|
| NER: ICD dict. | ||||
| Llama3.1:8b-instruct | 0.208 | 0.088 | 0.124 | 0.066 |
| Llama3-Med42-8B | 0.202 | 0.084 | 0.118 | 0.063 |
| Phi-3.5-mini-instruct | 0.211 | 0.093 | 0.129 | 0.069 |
| Mistral-Nemo-Instruct-2407 | 0.198 | 0.072 | 0.105 | 0.055 |
| Qwen2.5-7B-Instruct | 0.206 | 0.087 | 0.122 | 0.065 |
| NER: ICD dict. + RuCCoD | ||||
| Llama3.1:8b-instruct | 0.581 | 0.456 | 0.511 | 0.343 |
| Llama3-Med42-8B | 0.556 | 0.441 | 0.492 | 0.326 |
| Phi-3.5-mini-instruct | 0.543 | 0.450 | 0.492 | 0.326 |
| Mistral-Nemo-Instruct-2407 | 0.541 | 0.372 | 0.441 | 0.283 |
| Qwen2.5-7B-Instruct | 0.566 | 0.440 | 0.495 | 0.329 |
| NER+Linking: ICD dict. | ||||
| Llama3.1:8b-instruct | 0.071 | 0.067 | 0.069 | 0.036 |
| Llama3-Med42-8B | 0.058 | 0.063 | 0.060 | 0.031 |
| Phi-3.5-mini-instruct | 0.062 | 0.069 | 0.065 | 0.034 |
| Mistral-Nemo-Instruct-2407 | 0.066 | 0.056 | 0.060 | 0.031 |
| Qwen2.5-7B-Instruct | 0.065 | 0.065 | 0.065 | 0.033 |
| NER+Linking: ICD dict. + RuCCoD | ||||
| Llama3.1:8b-instruct | 0.272 | 0.264 | 0.268 | 0.155 |
| Llama3-Med42-8B | 0.235 | 0.261 | 0.247 | 0.141 |
| Phi-3.5-mini-instruct | 0.228 | 0.257 | 0.242 | 0.137 |
| Mistral-Nemo-Instruct-2407 | 0.247 | 0.215 | 0.230 | 0.130 |
| Qwen2.5-7B-Instruct | 0.244 | 0.246 | 0.245 | 0.140 |
| Code assignment: ICD dict. | ||||
| Llama3.1:8b-instruct | 0.379 | 0.363 | 0.371 | 0.228 |
| Llama3-Med42-8B | 0.310 | 0.345 | 0.327 | 0.195 |
| Phi-3.5-mini-instruct | 0.260 | 0.294 | 0.276 | 0.160 |
| Mistral-Nemo-Instruct-2407 | 0.413 | 0.360 | 0.385 | 0.238 |
| Qwen2.5-7B-Instruct | 0.401 | 0.411 | 0.406 | 0.255 |
| Code assignment: ICD dict. + RuCCoD | ||||
| Llama3.1:8b-instruct | 0.465 | 0.451 | 0.458 | 0.297 |
| Llama3-Med42-8B | 0.434 | 0.483 | 0.457 | 0.296 |
| Phi-3.5-mini-instruct | 0.409 | 0.458 | 0.432 | 0.276 |
| Mistral-Nemo-Instruct-2407 | 0.462 | 0.401 | 0.429 | 0.273 |
| Qwen2.5-7B-Instruct | 0.461 | 0.465 | 0.463 | 0.301 |
🔼 This table presents the performance of different Large Language Models (LLMs) with Retrieval-Augmented Generation (RAG) on three tasks related to ICD coding: Named Entity Recognition (NER), linking entities to ICD codes, and end-to-end entity linking. It shows precision, recall, F1-score, and accuracy for each LLM on each task, using different knowledge sources (ICD dictionary, RuCCOD dataset) for the RAG component.
read the caption
Table 9: Evaluation results for NER, Code assignment, and end-to-end entity linking task on RuCCoD for LLM+RAG pipeline.
| Model | Precision | Recall | F-score | Accuracy |
|---|---|---|---|---|
| NER: NEREL-BIO | ||||
| Llama3.1:8b-instruct | 0.100 | 0.042 | 0.059 | 0.030 |
| Llama3-Med42-8B | 0.104 | 0.043 | 0.060 | 0.031 |
| Phi-3.5-mini-instruct | 0.098 | 0.043 | 0.059 | 0.031 |
| Mistral-Nemo-Instruct-2407 | 0.115 | 0.044 | 0.063 | 0.033 |
| Qwen2.5-7B-Instruct | 0.099 | 0.043 | 0.060 | 0.031 |
| NER: RuCCoN | ||||
| Llama3.1:8b-instruct | 0.188 | 0.088 | 0.120 | 0.064 |
| Llama3-Med42-8B | 0.174 | 0.079 | 0.108 | 0.057 |
| Phi-3.5-mini-instruct | 0.172 | 0.085 | 0.114 | 0.060 |
| Mistral-Nemo-Instruct-2407 | 0.197 | 0.082 | 0.116 | 0.061 |
| Qwen2.5-7B-Instruct | 0.185 | 0.091 | 0.122 | 0.065 |
| NER+Linking: NEREL-BIO | ||||
| Llama3.1:8b-instruct | 0.023 | 0.020 | 0.021 | 0.011 |
| Llama3-Med42-8B | 0.018 | 0.019 | 0.018 | 0.009 |
| Phi-3.5-mini-instruct | 0.019 | 0.020 | 0.019 | 0.010 |
| Mistral-Nemo-Instruct-2407 | 0.025 | 0.020 | 0.022 | 0.011 |
| Qwen2.5-7B-Instruct | 0.021 | 0.020 | 0.020 | 0.010 |
| NER+Linking: RuCCoN | ||||
| Llama3.1:8b-instruct | 0.050 | 0.046 | 0.048 | 0.025 |
| Llama3-Med42-8B | 0.042 | 0.044 | 0.043 | 0.022 |
| Phi-3.5-mini-instruct | 0.038 | 0.041 | 0.040 | 0.020 |
| Mistral-Nemo-Instruct-2407 | 0.053 | 0.044 | 0.048 | 0.025 |
| Qwen2.5-7B-Instruct | 0.048 | 0.046 | 0.047 | 0.024 |
| Code assignment: NEREL-BIO | ||||
| Llama3.1:8b-instruct | 0.059 | 0.053 | 0.056 | 0.029 |
| Llama3-Med42-8B | 0.045 | 0.047 | 0.046 | 0.024 |
| Phi-3.5-mini-instruct | 0.046 | 0.049 | 0.047 | 0.024 |
| Mistral-Nemo-Instruct-2407 | 0.062 | 0.051 | 0.056 | 0.029 |
| Qwen2.5-7B-Instruct | 0.058 | 0.056 | 0.057 | 0.029 |
| Code assignment: RuCCoN | ||||
| Llama3.1:8b-instruct | 0.164 | 0.150 | 0.157 | 0.085 |
| Llama3-Med42-8B | 0.125 | 0.131 | 0.128 | 0.068 |
| Phi-3.5-mini-instruct | 0.125 | 0.134 | 0.129 | 0.069 |
| Mistral-Nemo-Instruct-2407 | 0.156 | 0.129 | 0.141 | 0.076 |
| Qwen2.5-7B-Instruct | 0.156 | 0.152 | 0.154 | 0.084 |
🔼 This table presents the performance of different Large Language Models (LLMs) with Retrieval Augmented Generation (RAG) on three tasks related to ICD coding: Named Entity Recognition (NER), NER+linking, and code assignment. The evaluation was performed on the RuCCoD dataset, using either NEREL-BIO or RuCCoN as vectorstores. The results show precision, recall, F-score, and accuracy for each model and task. This helps assess the efficacy of different LLMs and approaches in automating various stages of the ICD coding process.
read the caption
Table 10: Evaluation results for NER, Code assignment, and end-to-end entity linking task on RuCCoD for LLM+RAG pipeline using NEREL-BIO and RuCCoN for vectorstore.
| Model | Precision | Recall | F-score | Accuracy |
|---|---|---|---|---|
| NER: ICD dict. | ||||
| Llama3.1:8b-instruct | 0.208 | 0.088 | 0.124 | 0.066 |
| Llama3-Med42-8B | 0.202 | 0.084 | 0.118 | 0.063 |
| Phi-3.5-mini-instruct | 0.211 | 0.093 | 0.129 | 0.069 |
| Mistral-Nemo-Instruct-2407 | 0.198 | 0.072 | 0.105 | 0.055 |
| Qwen2.5-7B-Instruct | 0.206 | 0.087 | 0.122 | 0.065 |
| NER: ICD dict. + RuCCoD | ||||
| Llama3.1:8b-instruct | 0.581 | 0.456 | 0.511 | 0.343 |
| Llama3-Med42-8B | 0.556 | 0.441 | 0.492 | 0.326 |
| Phi-3.5-mini-instruct | 0.543 | 0.450 | 0.492 | 0.326 |
| Mistral-Nemo-Instruct-2407 | 0.541 | 0.372 | 0.441 | 0.283 |
| Qwen2.5-7B-Instruct | 0.566 | 0.440 | 0.495 | 0.329 |
| NER+Linking: ICD dict. | ||||
| Llama3.1:8b-instruct | 0.095 | 0.088 | 0.091 | 0.048 |
| Llama3-Med42-8B | 0.077 | 0.083 | 0.080 | 0.042 |
| Phi-3.5-mini-instruct | 0.083 | 0.092 | 0.087 | 0.046 |
| Mistral-Nemo-Instruct-2407 | 0.083 | 0.070 | 0.076 | 0.040 |
| Qwen2.5-7B-Instruct | 0.087 | 0.086 | 0.087 | 0.045 |
| NER+Linking: ICD dict. + RuCCoD | ||||
| Llama3.1:8b-instruct | 0.378 | 0.362 | 0.369 | 0.227 |
| Llama3-Med42-8B | 0.324 | 0.354 | 0.338 | 0.203 |
| Phi-3.5-mini-instruct | 0.323 | 0.357 | 0.339 | 0.204 |
| Mistral-Nemo-Instruct-2407 | 0.342 | 0.295 | 0.317 | 0.188 |
| Qwen2.5-7B-Instruct | 0.343 | 0.340 | 0.342 | 0.206 |
| Code assignment: ICD dict. | ||||
| Llama3.1:8b-instruct | 0.575 | 0.561 | 0.568 | 0.396 |
| Llama3-Med42-8B | 0.523 | 0.594 | 0.556 | 0.385 |
| Phi-3.5-mini-instruct | 0.437 | 0.510 | 0.471 | 0.308 |
| Mistral-Nemo-Instruct-2407 | 0.598 | 0.533 | 0.564 | 0.392 |
| Qwen2.5-7B-Instruct | 0.595 | 0.618 | 0.607 | 0.435 |
| Code assignment: ICD dict. + RuCCoD | ||||
| Llama3.1:8b-instruct | 0.701 | 0.684 | 0.692 | 0.529 |
| Llama3-Med42-8B | 0.644 | 0.720 | 0.680 | 0.515 |
| Phi-3.5-mini-instruct | 0.627 | 0.703 | 0.663 | 0.496 |
| Mistral-Nemo-Instruct-2407 | 0.691 | 0.605 | 0.645 | 0.476 |
| Qwen2.5-7B-Instruct | 0.700 | 0.704 | 0.702 | 0.541 |
🔼 This table presents the results of experiments evaluating the performance of different Large Language Models (LLMs) with Retrieval-Augmented Generation (RAG) on the RuCCoD dataset. The evaluation used a relaxed scoring approach, focusing on the NER (Named Entity Recognition), code assignment, and end-to-end entity linking tasks. The models are compared based on their precision, recall, F1-score, and accuracy, with results shown for various configurations, including the use of different dictionaries and datasets in the RAG pipeline. The results are analyzed to understand the effectiveness of each model in the relaxed setting.
read the caption
Table 11: Relaxed evaluation results for NER, Code assignment, and end-to-end entity linking task on RuCCoD for LLM+RAG pipeline.
| Model | Precision | Recall | F-score | Accuracy |
|---|---|---|---|---|
| NER: NEREL-BIO | ||||
| Llama3.1:8b-instruct-fp16 | 0.100 | 0.042 | 0.059 | 0.030 |
| Llama3-Med42-8B | 0.104 | 0.043 | 0.060 | 0.031 |
| Phi-3.5-mini-instruct | 0.098 | 0.043 | 0.059 | 0.031 |
| Mistral-Nemo-Instruct-2407 | 0.115 | 0.044 | 0.063 | 0.033 |
| Qwen2.5-7B-Instruct | 0.099 | 0.043 | 0.060 | 0.031 |
| NER: RuCCoN | ||||
| Llama3.1:8b-instruct-fp16 | 0.188 | 0.088 | 0.120 | 0.064 |
| Llama3-Med42-8B | 0.174 | 0.079 | 0.108 | 0.057 |
| Phi-3.5-mini-instruct | 0.172 | 0.085 | 0.114 | 0.060 |
| Mistral-Nemo-Instruct-2407 | 0.197 | 0.082 | 0.116 | 0.061 |
| Qwen2.5-7B-Instruct | 0.185 | 0.091 | 0.122 | 0.065 |
| NER+Linking: NEREL-BIO | ||||
| Llama3.1:8b-instruct | 0.033 | 0.029 | 0.031 | 0.016 |
| Llama3-Med42-8B | 0.024 | 0.025 | 0.025 | 0.013 |
| Phi-3.5-mini-instruct | 0.026 | 0.028 | 0.027 | 0.014 |
| Mistral-Nemo-Instruct-2407 | 0.033 | 0.027 | 0.030 | 0.015 |
| Qwen2.5-7B-Instruct | 0.030 | 0.029 | 0.030 | 0.015 |
| NER+Linking: RuCCoN | ||||
| Llama3.1:8b-instruct | 0.076 | 0.069 | 0.072 | 0.038 |
| Llama3-Med42-8B | 0.061 | 0.063 | 0.062 | 0.032 |
| Phi-3.5-mini-instruct | 0.060 | 0.064 | 0.062 | 0.032 |
| Mistral-Nemo-Instruct-2407 | 0.076 | 0.062 | 0.068 | 0.035 |
| Qwen2.5-7B-Instruct | 0.073 | 0.070 | 0.072 | 0.037 |
| Code assignment: NEREL-BIO | ||||
| Llama3.1:8b-instruct | 0.114 | 0.107 | 0.110 | 0.058 |
| Llama3-Med42-8B | 0.088 | 0.096 | 0.092 | 0.048 |
| Phi-3.5-mini-instruct | 0.098 | 0.110 | 0.104 | 0.055 |
| Mistral-Nemo-Instruct-2407 | 0.121 | 0.105 | 0.112 | 0.059 |
| Qwen2.5-7B-Instruct | 0.125 | 0.126 | 0.125 | 0.067 |
| Code assignment: RuCCoN | ||||
| Llama3.1:8b-instruct | 0.295 | 0.282 | 0.288 | 0.168 |
| Llama3-Med42-8B | 0.254 | 0.275 | 0.264 | 0.152 |
| Phi-3.5-mini-instruct | 0.248 | 0.273 | 0.260 | 0.149 |
| Mistral-Nemo-Instruct-2407 | 0.284 | 0.244 | 0.263 | 0.151 |
| Qwen2.5-7B-Instruct | 0.292 | 0.294 | 0.293 | 0.172 |
🔼 Table 12 presents the relaxed evaluation metrics for three tasks: Named Entity Recognition (NER), ICD code assignment, and end-to-end entity linking. The evaluation is performed on the RuCCoD dataset using the LLM+RAG pipeline. Specifically, the results showcase the performance of several large language models (LLMs) in these tasks when using either the NEREL-BIO or RuCCoN dataset as a vector store for retrieval augmented generation (RAG). The metrics presented include precision, recall, F-score, and accuracy, offering a comprehensive view of the models’ performance under relaxed evaluation conditions.
read the caption
Table 12: Relaxed evaluation results for NER, Code assignment, and end-to-end entity linking task on RuCCoD for LLM+RAG pipeline using NEREL-BIO and RuCCoN for vectorstore.
Full paper#